In a current research printed within the journal Nature Medicine, researchers assessed the coronavirus illness 2019 (COVID-19) vaccine effectiveness (VE) towards symptomatic infections with extreme acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants of concern (VOCs) and variants of curiosity (VOIs) by genetic mismatch/distance (GD) evaluation.
Vaccination is crucial to curtail transmission of SARS-CoV-2 and for mitigating COVID-19 severity. To date, a handful of SARS-CoV-2 vaccines are both within the early use part or have obtained approval for administration to the overall inhabitants. However, the safety conferred by vaccines is challenged by the emergence of novel genetic variants.
Study: Rapid analysis of COVID-19 vaccine effectiveness towards symptomatic an infection with SARS-CoV-2 variants by evaluation of genetic distance. Image Credit: Foxeel / Shutterstock
About the research
In the current research, researchers carried out GD evaluation to guage the VE of COVID-19 vaccines towards symptomatic SARS-CoV-2 infections based mostly on information from population-based epidemiological research.
The 4 vaccine platforms evaluated have been the messenger ribonucleic acid (mRNA)-based vaccines, protein subunit vaccines, viral vector-based vaccines, and inactivated vaccines, which have been obtained by 39, 5, 24, and 10 people, respectively. GDs have been calculated based mostly on the imply hamming distances on the receptor-binding domains (RBDs) of SARS-CoV-2 variants to the strains current within the vaccines.
VE was estimated by mixed-effects modeling by which GD was the important thing predictor and the confounders resembling age and the time elapsed put up the second vaccinations have been managed. VE and GD of the licensed vaccines have been in contrast, and subsequently, the group explored the impression of GD on immune safety conferred by vaccines. The VE values have been estimated towards SARS-CoV-2 VOCs resembling Alpha, Beta, Gamma, Delta, and Omicron (together with the BA.1 subvariant, the BA.1.1 subvariant, the BA.2 subvariant, and the BA.3 subvariant), and VOIs resembling Lambda and Mu.
Further, the VE-GD mannequin was assessed by validation information. VE information from 57 research and 23 research (variant-specific VE) have been used for mannequin coaching and validation, respectively. Further, SARS-CoV-2 variants have been predicted with their noticed VE not identified. The software of the estimated VE by the VE-GD mannequin was assessed in real-time towards the circulating SARS-CoV-2 variant in a selected geographical space, e.g., California.
Lastly, the group explored if region-wise vaccines may very well be developed and if they might match with the circulating SARS-CoV-2 pressure profiles, for which strains from 13 areas viz. the United Kingdom (UK), South Africa, Germany, India, Russia, Malaysia, Hong Kong, California, Japan, New York, Peru, Brazil, and Mexico have been analyzed.
Results
Among the vaccines assessed, VE estimates for the mRNA-based vaccines, protein subunit vaccines, viral vector-based vaccines, and inactivated vaccines have been 90%, 83%, 69%, and 60%, respectively. Of curiosity, the GD of the vaccines exhibited an reverse development, i.e., the least GD was noticed for the mRNA-based vaccines, whereas the opposite three vaccines exhibited bigger GD, and these variations may very well be because of the various intervals of vaccine analysis. The mRNA-based vaccine trials have been accomplished first throughout the interval when the SARS-CoV-2 inhabitants had comparatively increased homogeneity.
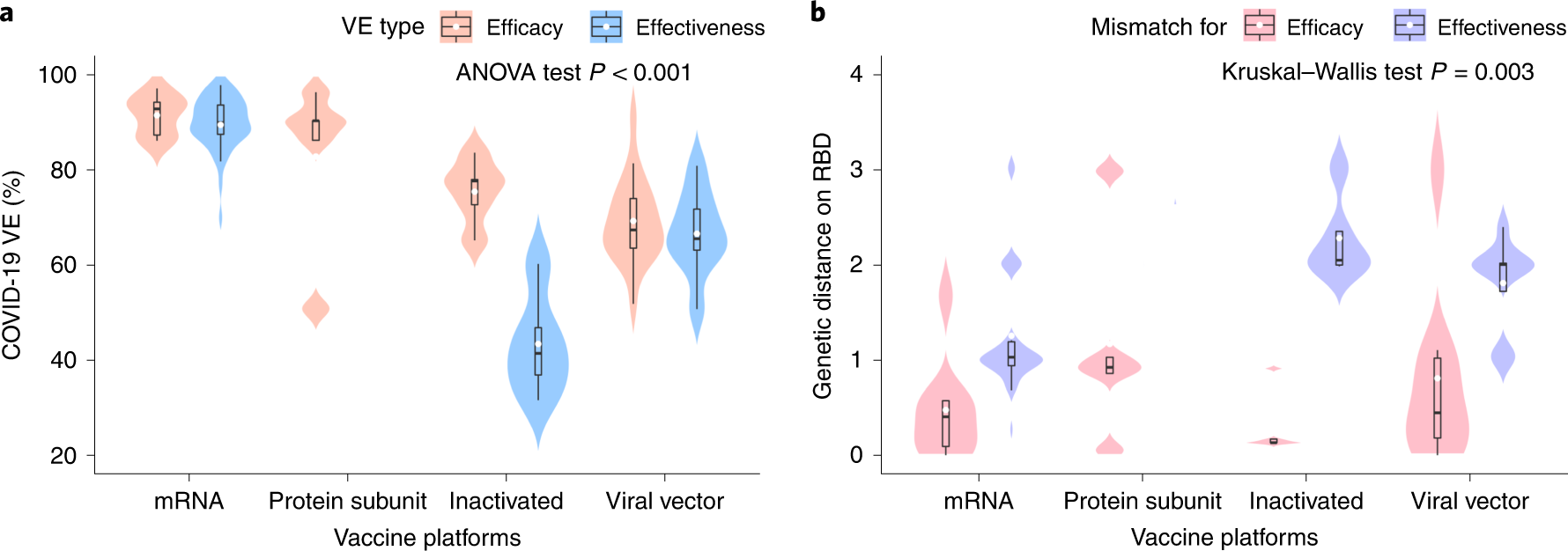 a, Distribution of the VE estimates for various platforms. The VE of mRNA and protein subunit vaccines are increased than different vaccines (two-sided ANOVA take a look at P = 2.2 × 10−14, n = 78). b, Distribution of genetic mismatch on RBD for various vaccine applied sciences. Genetic mismatch is the bottom for mRNA vaccines (two-sided Kruskal–Walls take a look at P = 0.003, n = 78). In the field plots, the center bar signifies the median; the white dot signifies the imply; and the boundaries are Q1 and Q3. Whiskers of the field plot are prolonged to Q3 + 1.5× interquartile vary (IQR) and Q1 − 1.5× IQR.
a, Distribution of the VE estimates for various platforms. The VE of mRNA and protein subunit vaccines are increased than different vaccines (two-sided ANOVA take a look at P = 2.2 × 10−14, n = 78). b, Distribution of genetic mismatch on RBD for various vaccine applied sciences. Genetic mismatch is the bottom for mRNA vaccines (two-sided Kruskal–Walls take a look at P = 0.003, n = 78). In the field plots, the center bar signifies the median; the white dot signifies the imply; and the boundaries are Q1 and Q3. Whiskers of the field plot are prolonged to Q3 + 1.5× interquartile vary (IQR) and Q1 − 1.5× IQR.
Over 86% and 88% of VE variations have been explainable based mostly on their GDs after the random results of the vaccine technological platforms and people of the vaccine merchandise (viz. mRNA-1273, BNT162b2, Ad26.COV2.S, AZD1222, CoronaVac and NVX-CoV2373), respectively have been managed. GD on the S RBD demonstrated probably the most profound impression on the safety conferred by vaccines, whereas no hyperlink was discovered between VE and GD of different SARS-CoV-2 proteins. With every S RBD substitution, VE would lower by 5.2%, 6.8%, 14.3%, and 15.8% for the mRNA-based vaccines, viral vector-based vaccines, protein subunit vaccines, and inactivated vaccines, respectively.
The S protein and N-terminal area (NTD) demonstrated weaker associations between VE and amino acid substitutions. In no GD was current, the VE (by the RBD space) for mRNA-based vaccines and protein subunit vaccines was estimated to be ~96%, whereas the VE for the viral vector-based vaccines and inactivated vaccines have been estimated to be lesser by 20.6% and 17.3%, respectively.
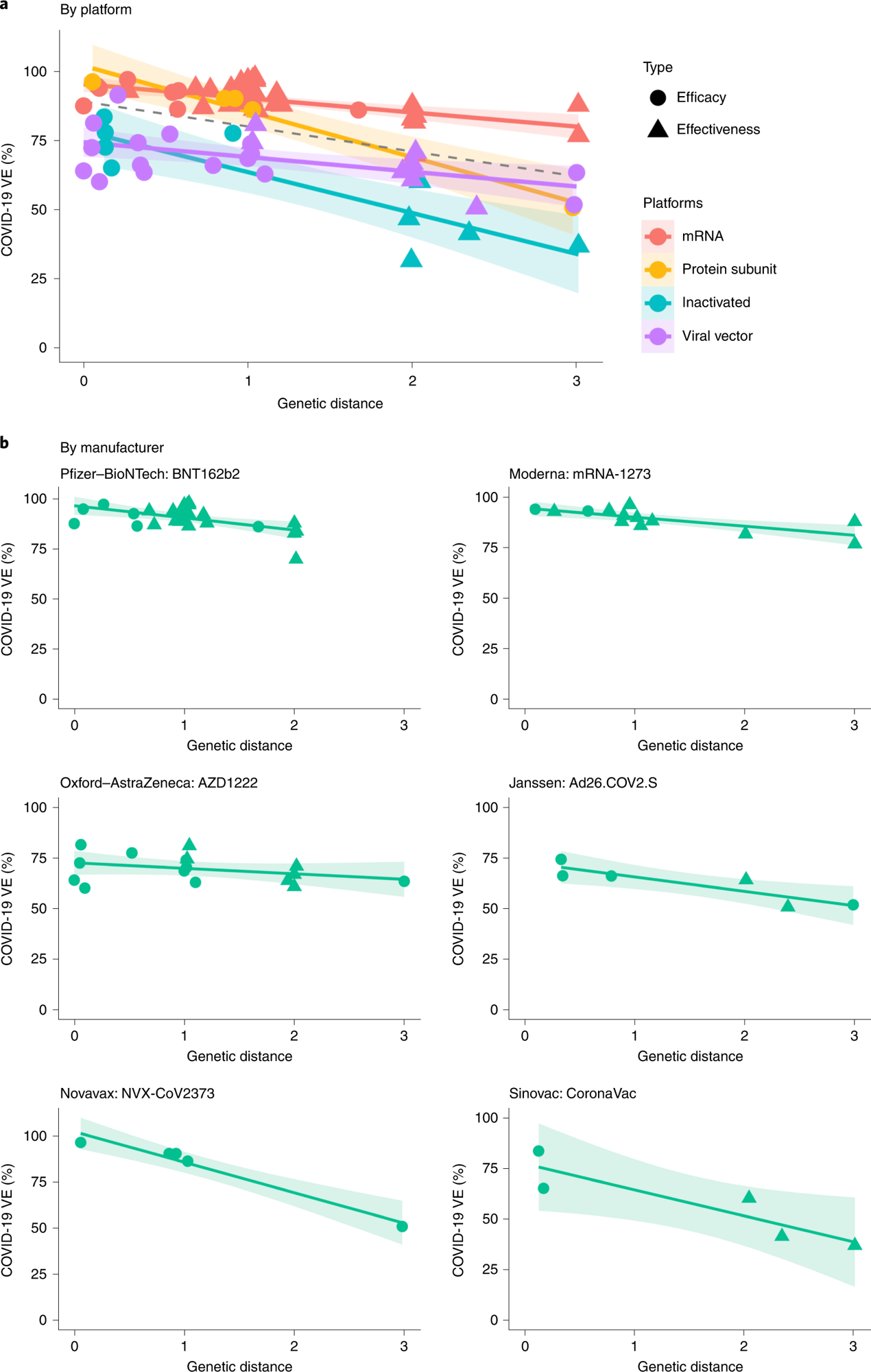
a, Negative linear relationships between VE and GD for various vaccine platforms (P = 0.038, R2 = 86.3%). The dashed line was fitted by all information factors. b, Negative linear relationship between VE and GD for every vaccine product (P = 0.006, R2 = 87.9%). The two-sided P worth was obtained from the mixed-effects mannequin. The coloured strains have been fitted by information factors of every platform. The shaded space signifies 95% CI.
Predicted VE estimates of the mRNA-based vaccines and the viral vector-based vaccines for Delta have been 82.8% and 61%, respectively, and the corresponding noticed VE values have been 83% and 67% for the BNT162b2 mRNA-based vaccine and the AZD1222 l vector-based vaccines, respectively. Likewise, predicted VE estimates for the BNT162b2 and mRNA-1273 vaccines have been 89.4% towards Alpha and 73.7% towards Beta and Gamma, much like the corresponding noticed VE values of 86% and 77%, respectively.
The predicted VE for the mRNA-1273 vaccine towards Omicron was 14%, much like the noticed VE of 13.9% in December 2021 in California. The findings have been indicative of the elevated validity of the VE-GD method. Predicted VE estimates for mRNA-based vaccines towards the Omicron subvariants have been between 11.9 % (Omicron BA.1) and 33.3% (Omicron BA.2).
The mannequin predicted that VE estimates for VOCs apart from Omicron and VOIs resembling Mu and Lambda can be >50% after three months of double mRNA-based vaccination; nevertheless, the VE estimates for inactivated vaccines towards SARS-CoV-2 infections (symptomatic) have been estimated to lower with the emergence of novel genetic variants of SARS-CoV-2.
The Omicron subvariants may very well be matched to all areas investigated besides Russia within the interval between January 2022 and February 2022; nevertheless, completely different subvariants have been predominant in numerous geographical areas. The findings indicated that updating vaccine formulations with just one variant could also be insufficient for matching the viral populations current throughout the globe.
Conclusion
Overall, the research findings highlighted the hyperlink between genetic mismatch of circulating SARS-CoV-2 variants and reported COVID-19 VE based mostly on the combination of epidemiological research and GD evaluation of SARS-CoV-2 variants. Furthermore, the findings indicated that GD values might considerably clarify VE alterations towards SARS-CoV-2 variants, and VE assessments towards evolving pathogens might help within the improvement of vaccines.