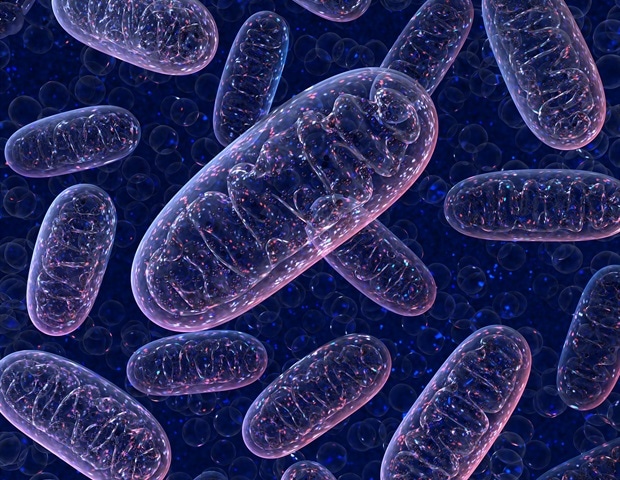
Mitochondria are compartments – so-called “organelles” — in our cells that provide the chemical energy supply we’d like to maneuver, think, and live. Chloroplasts are organelles in plants and algae that capture sunlight and perform photosynthesis. At a primary glance, they could look worlds apart. But a global team of researchers, led by the University of Bergen, have used data science and computational biology to point out that the identical “rules” have shaped how each organelles – and more – have evolved throughout life’s history.
Each forms of organelle were once independent organisms, with their very own full genomes. Billions of years ago, those organisms were captured and imprisoned by other cells – the ancestors of contemporary species. Since then, the organelles have lost most of their genomes, with only a handful of genes remaining in modern-day mitochondrial and chloroplast DNA. These remaining genes are essential for all times and necessary in lots of devastating diseases, but why they stay in organelle DNA – when so many others have been lost — has been debated for many years.
For a fresh perspective on this query, the scientists took a data-driven approach. They gathered data on all of the organelle DNA that has been sequenced across life. They then used modeling, biochemistry, and structural biology to represent a big selection of various hypotheses about gene retention as a set of numbers related to each gene. Using tools from data science and statistics, they asked which ideas could best explain the patterns of retained genes in the information that they had compiled – testing the outcomes with unseen data to envision their power.
Some clear patterns emerged from the modeling. Plenty of these genes encode subunits of larger cellular machines, that are assembled like a jigsaw. Genes for the pieces in the midst of the jigsaw are most definitely to remain in organelle DNA.”
Kostas Giannakis, postdoctoral researcher at Bergen and joint first creator on the paper
The team believes that it is because keeping local control over the production of such central subunits help the organelle quickly respond to vary – a version of the so-called “CoRR” model. Additionally they found support for other existing, debated, and latest ideas. For instance, if a gene product is hydrophobic – and hard to import to the organelle from outside – the information shows that it is commonly retained there. Genes which are themselves encoded using stronger-binding chemical groups are also more often retained – perhaps because they’re more robust in the cruel environment of the organelle.
“These different hypotheses have often been considered competing up to now,” says Iain Johnston, a professor at Bergen and leader of the team. “But actually no single mechanism can explain all of the observations – it takes a mixture. A strength of this unbiased, data-driven approach is that it will possibly show that a number of ideas are partly right, but none exclusively so – perhaps explaining the long debate on these topics.”
To their surprise, the team also found that their models trained to explain mitochondrial genes also predicted the retention of chloroplast genes, and vice versa. Additionally they found that the identical genetic features shaping mitochondrial and chloroplast DNA also appear to play a task within the evolution of other endosymbionts – organisms which have been more recently captured by other hosts, from algae to insects.
“That was a wow moment,” says Johnston. “We – and others – have had this concept that similar pressures might apply to the evolution of various organelles. But to see this universal, quantitative link – data from one organelle precisely predicting patterns in one other, and in more moderen endosymbionts – was really striking.”
The research is an element of a broader project funded by the European Research Council, and the team at the moment are working on a parallel query – how different organisms maintain the organelle genes that they do retain. Mutations in mitochondrial DNA may cause devastating inherited diseases; the team is using modeling, statistics, and experiments to explore how these mutations are handled in humans, plants, and more.
Source:
Journal reference:
Giannakis, K., et al. (2022) Evolutionary inference across eukaryotes identifies universal features shaping organelle gene retention. Cell Systems. doi.org/10.1016/j.cels.2022.08.007.